2022 Blogs, Blog, Featured
Pipelining GATK with WDL and Cromwell on AWS Cloud
Developed in the Data Sciences Platform at the Broad Institute, the Genome Analysis Toolkit (GATK) offers a wide variety of tools with a primary focus on variant discovery and genotyping. Relevance Lab is pleased to offer researchers the ability to run their GATK pipelines on AWS that was missing so far with our Genomics Cloud solution and a 1-click model.
GATK is making scientific research simpler for Genomics by providing best practices workflows and docker containers. The workflows are written in Workflow Description Language (WDL), a user-friendly scripting language maintained by the OpenWDL community. Cromwell is an open-source workflow execution engine that supports WDL as well as CWL, the Common Workflow Language, and can be run on a variety of different platforms, both local and cloud-based. RLCatalyst Research Gateway added support for the Cromwell engine that enables researchers to run any popular workflows on AWS seamlessly. Some of the popular workflows that are available for a quick start are the following:
- Mitochondrial short variant discovery (SNVs + Indels)
- Somatic short variant discovery (SNVs + Indels)
- Somatic copy number variant discovery (CNVs)
- Germline short variant discovery (SNPs + Indels)
- Germline copy number variant discovery (CNVs)
- RNAseq short variant discovery (SNPs + Indels)
The figure below shows the building block of this solution on AWS Cloud.
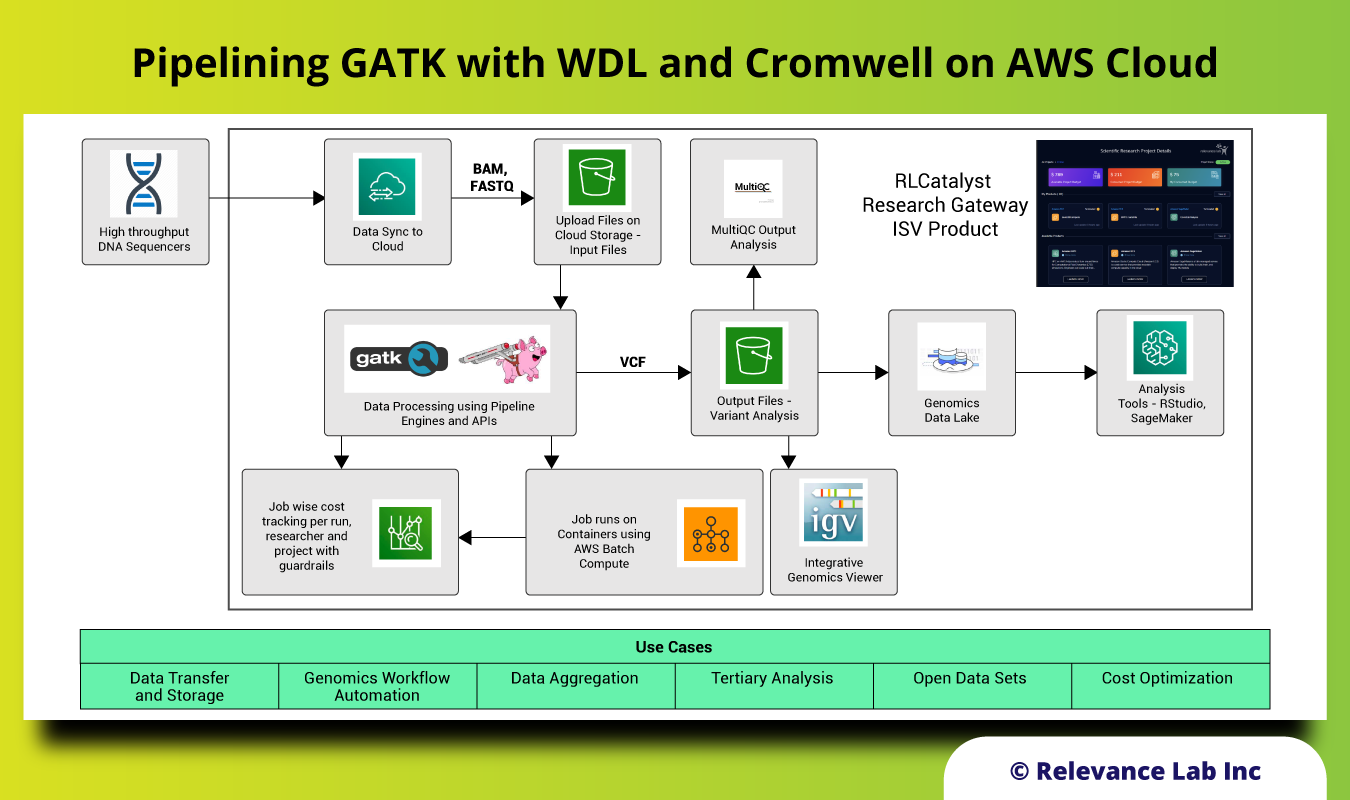
Steps for running GATK with WDL and Cromwell on AWS Cloud
| Steps | Details | Time Taken |
|---|---|---|
| 1. | Log into RLCatalyst Research Gateway as a Principal Investigator or Researcher profile. Select the project for running Genomics Pipelines, and first time create a new Cromwell Advanced Product. | 5 min |
| 2. | Select the Input Data location, output data location, pipeline to run (from GATK), and provide parameters (input.json). Default parameters are already suggested for the use of AWS Batch with Spot instances and all other AWS complexities, abstracted from the end-user, for simplicity. | 5 min to provision new Cromwell Server on AWS with AWS Batch setup completed with 1-Click |
| 3. | Execute Pipeline (using UI interface or by SSH into Head-node) on Cromwell Server. There is ability to run the new pipelines, monitor status, and review outputs from within the Portal UI. | Pipelines can take some time to run depending on size of data and complexity |
| 4. | View outputs of the Pipeline in Outputs S3 bucket from within the Portal. Use specialized tools like MultiQC, Integrative Genomics Viewer (IGV), and RStudio for further analysis. | 5 min |
| 5. | All costs related to User, Product, and Pipelines are automatically tagged and can be viewed in the budgets screen to know the cloud spend for pipeline execution that consists of all resources, including AWS Batch HPC instances dynamically provisioned. Once the pipelines are executed, the existing Cromwell Server can be stopped or terminated to reduce ongoing costs. | 5 min |
The figure below shows the ability to select Cromwell Advanced to provision and run any pipeline.
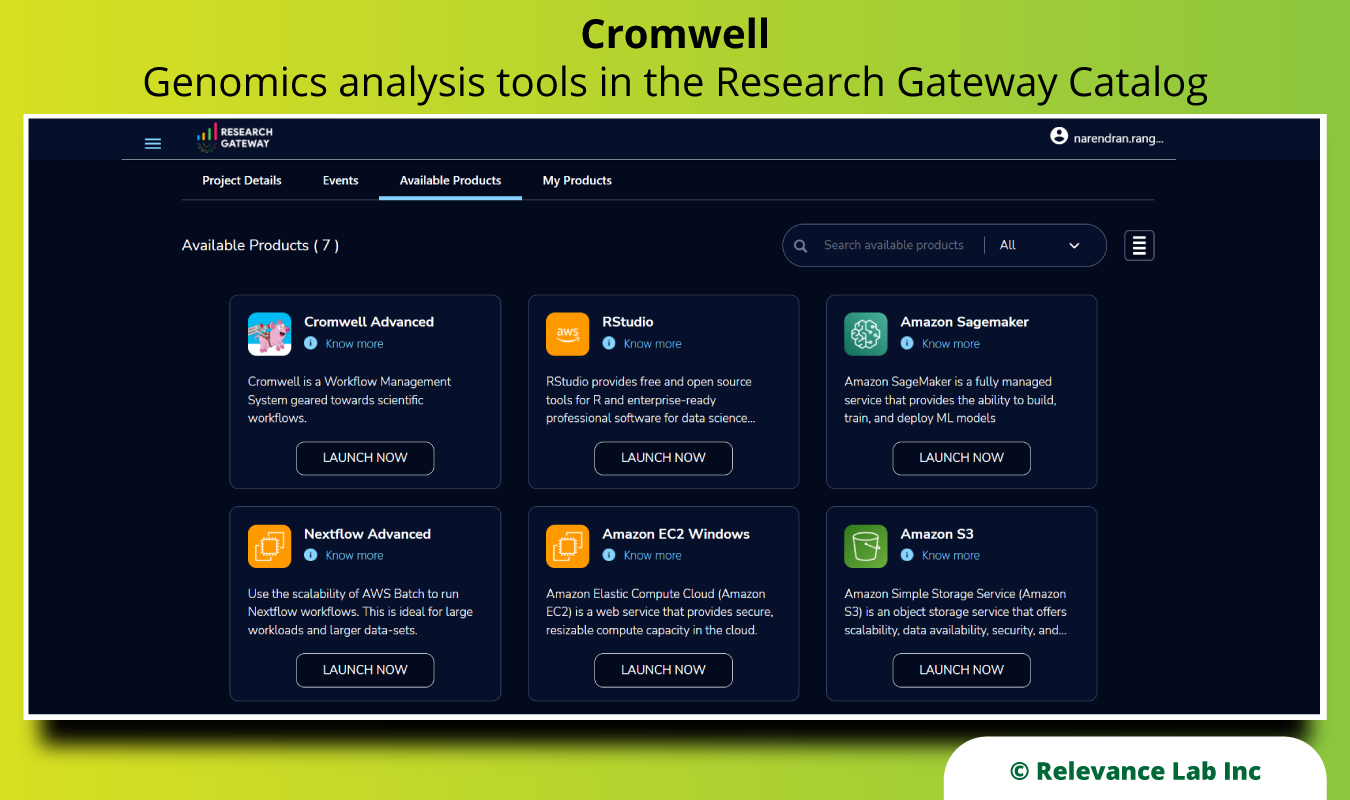
The following picture shows the architecture of Cromwell on AWS.
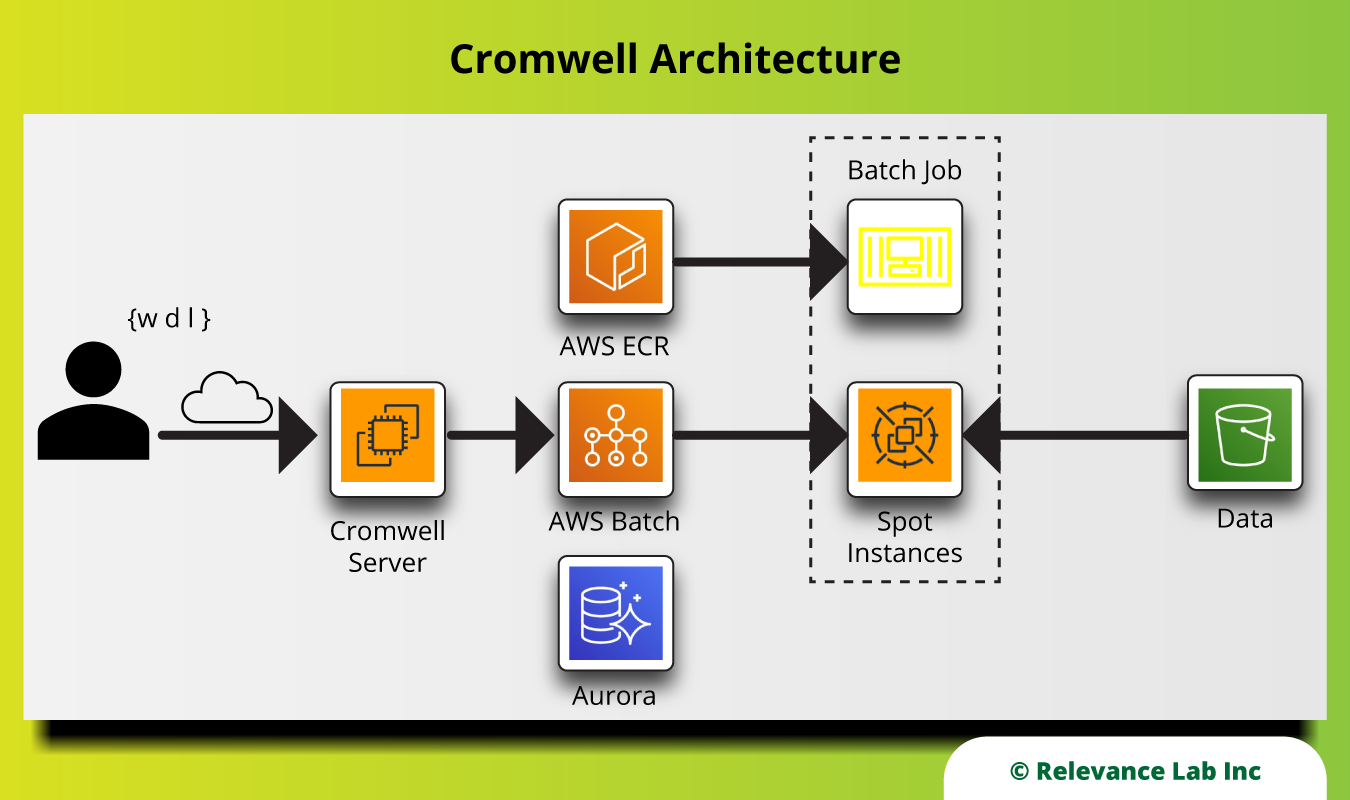
Summary
GATK community is constantly striving to make Genomics Research in the cloud simpler. So far, the support for AWS Cloud was still missing and was a key ask from multiple online research communities. Relevance Lab, in partnership with AWS, has addressed this need with their Genomics Cloud solution to make scientific research frictionless.
To know more about how you can start your GATK pipelines with WDL and Cromwell on the AWS Cloud in just 30 minutes using our solution at https://research.rlcatalyst.com, feel free to write to marketing@relevancelab.com
References
Accelerating Analytics for the Future of Genomics
Cromwell on AWS
Leveraging AWS HPC for Accelerating Scientific Research on Cloud
Cromwell Documentation
Artificial Intelligence, Machine Learning and Genomics
Accelerating Genomics and High Performance Computing on AWS with Relevance Lab Research Gateway Solution


